
Unlocking Statistical Power: Mastering Advanced Bayesian Modeling with Stan
Discover how Stan's flexible framework empowers researchers to build sophisticated statistical models for complex real-world data analysis
Key Insights into Advanced Bayesian Modeling with Stan
- Hierarchical modeling capabilities enable sophisticated analysis of nested data structures with partial pooling that balances individual variation and group-level patterns
- Efficient MCMC sampling through the No-U-Turn Sampler (NUTS) algorithm makes complex posterior inference computationally feasible
- Flexible model specification allows for implementation of custom likelihood functions, prior distributions, and complex model structures
Understanding Stan as a Bayesian Modeling Framework
Stan is a state-of-the-art probabilistic programming language specifically designed for Bayesian statistical modeling and computation. It provides a flexible framework that allows researchers and data scientists to specify complex statistical models and perform Bayesian inference using advanced Markov Chain Monte Carlo (MCMC) methods, particularly Hamiltonian Monte Carlo (HMC) with the No-U-Turn Sampler (NUTS).
What sets Stan apart from other statistical tools is its ability to handle highly complex model structures while maintaining computational efficiency. This makes it particularly valuable for advanced applications across various domains including social sciences, ecology, epidemiology, sports analytics, economics, and many other fields requiring sophisticated statistical analysis.
Core Components of the Stan Ecosystem
The Stan ecosystem consists of several integrated components:
- A modeling language for specifying probability models
- Inference algorithms (primarily HMC and NUTS) for sampling from posterior distributions
- Interfaces with popular programming languages (R, Python, Julia, Stata, etc.)
- Tools for model checking, validation, and comparison
- An active community developing extensions and applications
Stan's interfaces with other programming environments (such as RStan for R or PyStan for Python) allow users to integrate Stan models seamlessly into their existing workflows, making it accessible to researchers from diverse backgrounds.
Hierarchical Modeling: A Cornerstone of Advanced Bayesian Analysis
Hierarchical (or multilevel) modeling represents one of Stan's most powerful capabilities for advanced Bayesian analysis. These models are particularly valuable when data has a nested or grouped structure, such as students within schools, patients within hospitals, or measurements repeated over time.
The Concept of Partial Pooling
A key advantage of hierarchical models in Stan is the implementation of partial pooling, which strikes a balance between complete pooling (treating all groups as identical) and no pooling (analyzing each group independently). With partial pooling, the degree to which information is shared across groups is determined by the data itself:
- When groups show similar patterns, stronger pooling occurs (reflected in low hierarchical variance)
- When groups exhibit substantial differences, weaker pooling occurs (reflected in higher hierarchical variance)
- This adaptive approach improves estimation, especially for groups with limited data
Example: The Eight Schools Problem
A classic example demonstrating hierarchical modeling in Stan is the "Eight Schools" problem, where the goal is to estimate the effects of coaching programs on test scores across eight different schools. Each school has its own treatment effect, but these effects are related through a population distribution. Stan efficiently handles this multilevel structure, allowing for more accurate and nuanced estimates than traditional methods.
In this tutorial video, you can learn about hierarchical Bayesian modeling with Stan and techniques for optimizing Stan code for more efficient model fitting and inference.
Advanced Model Types and Techniques in Stan
Stan supports a wide range of advanced modeling techniques beyond simple linear models. These sophisticated approaches allow researchers to capture complex patterns and relationships in their data.
Complex Model Structures
- Non-linear relationships - When relationships between variables cannot be adequately captured by linear functions
- Time series models - For data collected over time with temporal dependencies
- Spatial models - For data with geographic dependencies
- Mixture models - When data comes from multiple underlying distributions
- Gaussian processes - For flexible, non-parametric function estimation
- Latent variable models - For incorporating unobserved variables that influence observed data
Multivariate Priors for Hierarchical Models
In advanced hierarchical modeling, specifying appropriate priors for group-level parameters becomes crucial. Stan provides flexible options for specifying multivariate priors, allowing for correlation structures among parameters. This capability is particularly important when modeling complex systems where various factors may be interrelated.
| Model Type | Key Features | Common Applications | Stan Implementation Complexity |
|---|---|---|---|
| Hierarchical Linear Models | Varying intercepts/slopes, partial pooling | Educational research, sports analytics | Moderate |
| Hierarchical Logistic Regression | Binary outcomes with grouped structure | Medical trials, voting behavior | Moderate to High |
| Time Series Models | Temporal dependencies, seasonality | Economic forecasting, environmental monitoring | High |
| Spatial Models | Geographic dependencies, autocorrelation | Epidemiology, ecology, real estate | Very High |
| Gaussian Process Models | Flexible function estimation | Machine learning, nonlinear dynamics | Very High |
Stan Implementation and Workflow
Implementing advanced Bayesian models in Stan follows a structured workflow that helps ensure reliable results and valid inferences.
The Bayesian Modeling Workflow in Stan
1. Model Specification
Stan uses a domain-specific language for model specification, with blocks for data, parameters, transformed parameters, model (likelihood and priors), and generated quantities. This structured approach ensures clarity and reproducibility.
2. Efficient Parameterization
Advanced modeling in Stan often requires careful parameterization to ensure efficient sampling. Non-centered parameterization (also called the "Matt trick") is particularly useful for hierarchical models to avoid issues like divergent transitions.
3. Model Validation and Diagnostics
Stan provides various tools for assessing model convergence and fit, including checks for divergent transitions, effective sample size, and Rhat statistics. Posterior predictive checks help validate whether the model can generate data similar to observed data.
4. Model Comparison and Selection
Information criteria like WAIC (Widely Applicable Information Criterion) and LOO-CV (Leave-One-Out Cross-Validation) help compare alternative models and select the most appropriate one for the data.
Example Code: Hierarchical Model in Stan
data {
int<lower=0> N; // number of observations
int<lower=0> J; // number of groups
vector[N] y; // outcome variable
int<lower=1,upper=J> id[N]; // group indicator
}
parameters {
real mu; // population mean
real<lower=0> tau; // population standard deviation
vector[J] eta; // standardized group-level effects
}
transformed parameters {
vector[J] theta; // group-level effects
theta = mu + tau * eta; // non-centered parameterization
}
model {
// Priors
mu ~ normal(0, 5);
tau ~ cauchy(0, 2.5);
eta ~ normal(0, 1);
// Likelihood
y ~ normal(theta[id], sigma);
}Performance and Comparative Analysis
When considering advanced Bayesian modeling approaches, it's valuable to understand how Stan compares to other frameworks and what performance characteristics to expect.
This radar chart compares Stan with other popular Bayesian modeling frameworks across key dimensions. Stan particularly excels in flexibility, handling complex hierarchical structures, and providing robust model diagnostics, making it especially suitable for advanced modeling applications.
Key Capabilities and Applications of Stan
This mindmap illustrates the extensive capabilities and applications of Stan for advanced Bayesian modeling. The framework supports diverse model types, statistical techniques, and application domains, with multiple implementation interfaces making it accessible across different programming environments.
Interface Options and Extensions
Stan's flexibility is enhanced by various interfaces and extension packages that make it more accessible and powerful for specific applications.
Stan Interfaces
Stan can be accessed through multiple interfaces, accommodating users from different programming backgrounds:
- RStan - The R interface to Stan, allowing Stan models to be specified and run from within R
- PyStan - The Python interface, bringing Stan's capabilities to Python users
- CmdStan - A command-line interface, useful for server environments or batch processing
- Stan.jl - An interface for Julia users
- StataStan - Allows Stata users to utilize Stan's capabilities
High-Level Packages and Extensions
Several packages build on Stan to provide higher-level interfaces for specific modeling needs:
brms (Bayesian Regression Models using Stan)
The brms package provides an accessible interface for fitting Bayesian generalized (non-)linear multilevel models using Stan. It uses familiar R formula syntax, making it easier for R users to specify complex models without writing Stan code directly.
rstanarm
This package provides a collection of pre-compiled Stan models for common regression applications, allowing users to fit Bayesian models using familiar R syntax while leveraging Stan's computational efficiency.
Torsten
An extension for pharmacometrics and pharmacokinetic/pharmacodynamic (PK/PD) modeling, Torsten provides specialized functions for modeling drug absorption, distribution, metabolism, and excretion.
Visual Gallery: Stan in Action
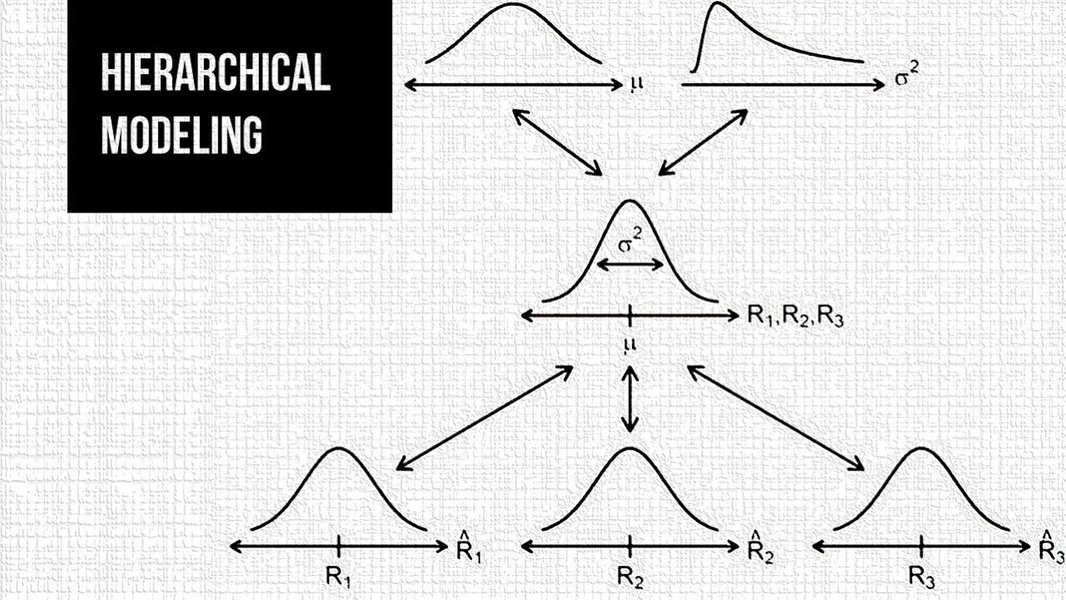
Hierarchical Bayesian modeling in Stan allows researchers to analyze complex nested data structures with elegant mathematical frameworks.

Computational modeling tutorials help users understand how to implement advanced statistical techniques in Stan.
Frequently Asked Questions
References
- Multivariate Priors for Hierarchical Models - Stan User's Guide
- Hierarchical Logistic Regression - Stan User's Guide
- Hierarchical models in Stan - Occasional Divergences
- Hierarchical models with RStan (Part 1) - Biology for Fun
- Advanced Bayesian Multilevel Modeling with the R Package brms - The R Journal
- Courses in Bayesian statistical modeling with Stan - Michael Betancourt
- Statistical Modeling Workshop Materials - GitHub Repository
Recommended Explorations
Last updated April 8, 2025