
Unraveling the Flavivirus Genome: A Blueprint for Viral Ingenuity
Explore the precise architecture and functional significance of the flavivirus genetic material, from its compact size to its critical protein-coding regions.
Key Insights into the Flavivirus Genome
- Single-Stranded Positive-Sense RNA: Flaviviruses possess a single-stranded, positive-sense RNA genome, enabling it to act directly as messenger RNA (mRNA) for protein synthesis upon entering a host cell.
- Typical Size Range: The average flavivirus genome spans approximately 9.7 to 12 kilobases (kb), with a highly consistent length of 10 to 11 kb across most well-studied species.
- Single Open Reading Frame (ORF): The genome contains one long ORF that translates into a large polyprotein, which is subsequently cleaved into three structural and seven nonstructural proteins, all essential for viral replication and assembly.
The Compact Blueprint: Flavivirus Genome Size and Structure
Flaviviruses, responsible for widespread diseases like dengue, Zika, and West Nile fever, encapsulate their genetic information within a highly efficient and compact genome. This genome is a single-stranded, positive-sense RNA molecule, meaning it functions directly as mRNA, allowing the host cell's machinery to translate it immediately into viral proteins upon infection. Its non-segmented nature, consisting of one continuous RNA strand, is a hallmark characteristic of these viruses.
Precise Dimensions: Understanding the Genome Length
The typical genome size of flaviviruses consistently falls within a narrow range. Most well-characterized flaviviruses have genomes approximately 10 to 11 kilobases (kb) in length. While some sources indicate a broader range from 9.7 kb to 12 kb, the 10-11 kb average represents the standard for the majority of species within the genus *Flavivirus*. Each kilobase signifies 1,000 nucleotides, highlighting the precise and relatively small size of this viral blueprint. This compact size facilitates efficient replication and packaging within the viral particle, contributing to the virus's ability to spread effectively.
Structural Elements: Untranslated Regions (UTRs)
The flavivirus genome is flanked by two crucial non-coding regions: the 5' untranslated region (UTR) at the beginning and the 3' UTR at the end. These regions, while not encoding proteins, are indispensable for viral replication and translation regulation. The 5' UTR is relatively short, typically around 100 nucleotides, and contains highly conserved sequences and intricate secondary structures vital for initiating translation and the subsequent replication process. In contrast, the 3' UTR is longer, ranging from 400 to 700 nucleotides, and features conserved RNA secondary structures critical for genome replication and stability. A notable feature is the absence of a poly-A tail at the 3' end, a common characteristic of eukaryotic mRNAs; instead, the 3' end forms a loop structure. The dynamic interaction between the 5' and 3' UTRs, facilitating a pseudo-circularization of the genome, is paramount for efficient RNA replication.
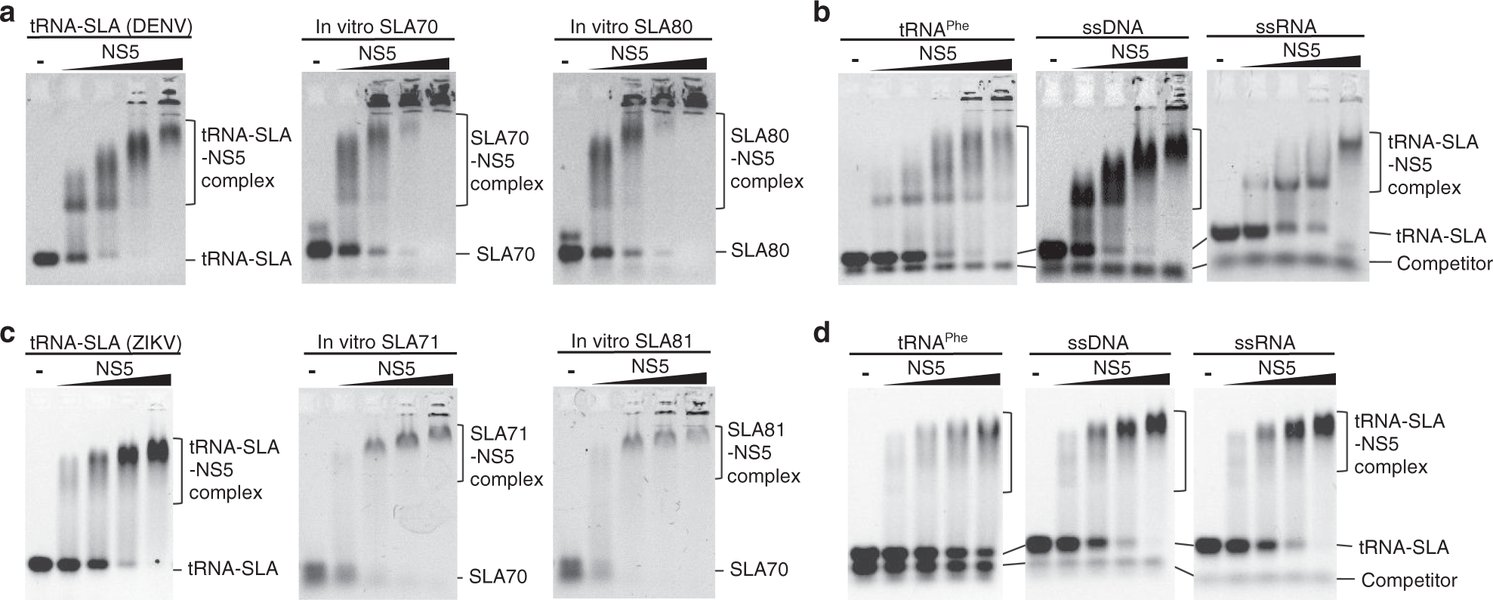
An illustration showing the crucial secondary structures within the flavivirus RNA, particularly in the untranslated regions, which are essential for its replication and function.
The Polyprotein Cascade: Encoding Viral Functions
Central to the flavivirus genome is a single, long open reading frame (ORF) that directs the synthesis of a large polyprotein. This polyprotein is then meticulously cleaved by viral and host proteases into individual, functional proteins. These proteins are categorized into two main groups: structural and nonstructural.
Structural Proteins: Building the Virion
The structural proteins are located at the 5' end of the coding region and are responsible for forming the viral particle (virion). There are three main structural proteins:
- Capsid (C) protein: Encapsulates and protects the viral RNA genome within the nucleocapsid.
- Precursor Membrane (prM) protein: A precursor that is cleaved into the mature membrane (M) protein during viral maturation. The prM protein plays a role in virion assembly and protects the Envelope protein from premature fusion.
- Envelope (E) protein: The most prominent surface protein, essential for viral entry into host cells. It mediates receptor binding and subsequent membrane fusion, allowing the viral genome to enter the cytoplasm. The E protein is also a primary target for neutralizing antibodies.
Nonstructural Proteins: Orchestrating Replication and Immune Evasion
The nonstructural proteins are located at the 3' end of the coding region and are involved in various crucial processes, including viral RNA replication, assembly of the replication complex, and evasion of the host's immune response. There are typically seven nonstructural proteins:
- NS1: Involved in RNA replication and immune evasion, existing as both a membrane-associated protein and a secreted form.
- NS2A: A small transmembrane protein involved in viral assembly and inhibition of host antiviral responses.
- NS2B: A cofactor for the NS3 protease, critical for polyprotein processing.
- NS3: A multifunctional enzyme possessing protease, helicase, and nucleoside triphosphatase (NTPase) activities, all vital for replication.
- NS4A: A transmembrane protein that induces membrane rearrangements, forming the replication compartments.
- NS4B: Another transmembrane protein involved in replication complex formation and antagonism of host innate immunity.
- NS5: The largest and most conserved nonstructural protein, functioning as the RNA-dependent RNA polymerase (RdRp) responsible for synthesizing new viral RNA genomes, and also possessing methyltransferase activity crucial for 5' cap formation.
Comparative Genome Features
To further illustrate the distinctive features of the flavivirus genome, here is a comparison of key aspects:
| Feature | Description | Significance |
|---|---|---|
| Genome Type | Single-stranded, Positive-sense RNA (+ssRNA) | Directly serves as mRNA, facilitating immediate protein synthesis upon infection. |
| Genome Size | Typically 10-11 kb (ranging 9.7-12 kb) | Compact size enables efficient replication and packaging within the virion. |
| Segmentation | Non-segmented (single linear molecule) | Ensures all genetic information is contained on one continuous strand. |
| 5' End Structure | Type 1 cap (m7GpppAmp) | Mimics host mRNA cap, essential for translation initiation and stability. |
| 3' End Structure | Lacks poly-A tail; forms conserved loop structures | Critical for replication efficiency and genome stability, interacting with the 5' UTR. |
| Coding Strategy | Single Open Reading Frame (ORF) | Encodes a polyprotein subsequently cleaved into individual proteins. |
| Proteins Encoded | 3 Structural (C, prM, E) and 7 Nonstructural (NS1-NS5) | Structural proteins form the virion; nonstructural proteins manage replication, assembly, and immune evasion. |
Visualizing the Flavivirus Genome's Functional Domains
To further understand the complex interplay of functions within the flavivirus genome, we can visualize its organization and the relative importance of its various components using a radar chart. This chart provides an opinionated analysis of how different elements contribute to the virus's lifecycle and pathogenicity, based on their known roles.
This radar chart illustrates the perceived contribution of different genome aspects to the overall flavivirus biology. "Genome Size (kb)" represents the typical length, with a score of 10.5 reflecting the average 10-11 kb range. "5' UTR Functionality" and "3' UTR Functionality" reflect the critical regulatory roles of these non-coding regions in replication and translation, with high scores indicating their importance despite their small size. "Structural Protein Importance" and "Nonstructural Protein Importance" highlight the essential roles of the encoded proteins, with nonstructural proteins often having a slightly higher score due to their direct involvement in replication machinery and immune evasion. Finally, "Replication Efficiency" encapsulates how well the entire genome structure supports rapid viral proliferation within the host. The hypothetical high virulence variant demonstrates how slight enhancements in these functional areas could contribute to increased viral fitness.
The Replication Cycle: A Deeper Dive
Understanding the flavivirus genome is incomplete without appreciating its role in the viral replication cycle. The positive-sense RNA genome acts as both the genetic material for progeny viruses and the direct template for immediate protein synthesis. This unique characteristic allows for rapid viral propagation upon entry into a susceptible host cell. The viral replication machinery, primarily composed of the nonstructural proteins, orchestrates the intricate steps of RNA replication on modified host cell membranes, particularly the rough endoplasmic reticulum. This process culminates in the production of new viral genomes and subsequent assembly into mature virions.
An animation illustrating the detailed replication cycle of the Dengue virus, a representative flavivirus. This video provides a visual understanding of how the flavivirus genome is utilized to replicate and assemble new viral particles within a host cell, showing the crucial steps from entry to release.
The replication cycle begins with the attachment of the Envelope (E) protein to host cell receptors, followed by receptor-mediated endocytosis. Once inside the endosome, the acidic environment triggers a conformational change in the E protein, leading to fusion of the viral and endosomal membranes. This releases the viral nucleocapsid into the cytoplasm, where the positive-sense RNA genome is uncoated. The genome then directly serves as mRNA for the synthesis of the polyprotein by host ribosomes. This polyprotein is subsequently cleaved into the individual structural and nonstructural proteins. The nonstructural proteins, particularly NS3 and NS5, form the core of the replication complex, which synthesizes a negative-sense RNA intermediate. This intermediate then serves as a template for the production of numerous positive-sense progeny genomes. These new genomes are then packaged into newly assembled virions, which bud into the endoplasmic reticulum and mature through the Golgi apparatus before being released from the cell.
Understanding Flavivirus Genome Organization
The flavivirus genome is a masterclass in compact genetic organization. Its single open reading frame and precise arrangement of structural and nonstructural genes, flanked by regulatory UTRs, represent an optimized blueprint for viral survival and propagation. The mindmap below illustrates this intricate architecture, highlighting the key components and their interconnections within the genome.
(9.7 - 12 kb range)"] id6["5' End Features"] id7["Type 1 Cap (m7GpppAmp)"] id8["5' UTR (~100 nt)"] id9["Highly conserved"] id10["Essential for translation & replication"] id11["Coding Region (Single ORF)"] id12["Polyprotein"] id13["Cleaved by proteases"] id14["Structural Proteins (5' end)"] id15["C (Capsid)"] id16["prM (Precursor Membrane)"] id17["E (Envelope)"] id18["Nonstructural Proteins (3' end)"] id19["NS1"] id20["NS2A"] id21["NS2B (Cofactor for NS3)"] id22["NS3 (Protease, Helicase)"] id23["NS4A"] id24["NS4B"] id25["NS5 (RdRp, Methyltransferase)"] id26["3' End Features"] id27["3' UTR (~400-700 nt)"] id28["Conserved RNA structures"] id29["Essential for replication & stability"] id30["No Poly-A Tail"] id31["Loop structure"]
This mindmap graphically represents the hierarchical organization of the flavivirus genome. Starting from the root, it branches out to key aspects like genome type and size, then delves into the specific features of the 5' and 3' ends, emphasizing the critical untranslated regions. The central branch details the single open reading frame, which gives rise to the polyprotein, subsequently cleaved into the essential structural and nonstructural proteins, each with its unique role in the viral life cycle. This visual aid simplifies the understanding of how various genomic elements contribute to the overall functionality of the flavivirus.
Frequently Asked Questions (FAQ)
Conclusion
The flavivirus genome, a single-stranded, positive-sense RNA molecule typically 10-11 kb in length, exemplifies evolutionary efficiency. Its compact design and singular open reading frame are meticulously organized to produce all the necessary structural and nonstructural proteins for replication and host interaction. The critical untranslated regions at both ends, while non-coding, govern the genome's translation and replication, enabling the virus to hijack host machinery effectively. This intricate genetic blueprint underpins the pathogenesis of significant global diseases, making its detailed understanding paramount for developing effective antiviral strategies and vaccines.
Recommended Further Exploration
- How do flaviviruses replicate their RNA genome inside host cells?
- What are the specific functions of each nonstructural protein in flavivirus replication?
- How do flaviviruses evade the host's immune system?
- What are the main challenges in developing vaccines against flaviviruses like Dengue or Zika?