
Unlocking the Genomic Code: A Deep Dive into the "3 Genomic Numbers"
Exploring the mathematical architecture that may govern the human genome's structure and function.

- 1.Key Insights into Genomic Numerical Architecture
- 2.The Foundation: Chargaff's Rules and the Constant 1
- 3.The Constant 2: Balancing Codon Frequencies
- 4.The Golden Ratio (Φ): Fine-Tuning Global Densities
- 5.The Genomic Meta-Architecture: A Universal Checksum
- 6.Perspectives and Consensus
- 7.The Interplay of Form and Substance
- 8.Implications for Future Research
- 9.Frequently Asked Questions
- 10.Conclusion
- 11.Recommended Further Exploration
- 12.Referenced Search Results
Key Insights into Genomic Numerical Architecture
- The "3 Genomic Numbers" Theory: Jean-Claude Perez proposes that the human genome is governed by three fundamental constants—1, 2, and the Golden Ratio (Φ ≈ 1.618)—each playing a distinct role in its numerical organization.
- Chargaff's Parity and the Number 1: The constant 1 is linked to Chargaff's first parity rule, ensuring equal proportions of certain nucleotide bases (A=T, G=C) in DNA, balancing atomic mass between strands.
- Codon Frequency Balancing with Number 2: The number 2 is suggested to optimize codon frequency distributions within the Universal Genetic Code, fostering a fractal-like language resilient to mutations.
- Golden Ratio's Fine-Tuning: Φ is posited to fine-tune global densities, appearing in codon usage biases, GC-content, and fractal repetitions, creating a mathematical equilibrium between informational "Form" and structural "Substance."
- Mainstream vs. Perez's Framework: While Chargaff's rules are foundational in genomics, Perez's comprehensive "3 Genomic Numbers" framework is considered non-mainstream and has not achieved broad independent replication within the scientific community.
The intricate design of the human genome has long captivated scientists, leading to various theories attempting to explain its underlying structure and function. Among these, Jean-Claude Perez's "3 Genomic Numbers" framework stands out, proposing a deterministic, self-designed numerical architecture for DNA. This theory suggests that the human genome is not merely a random sequence but an optimized system governed by specific mathematical constants: 1, 2, and the Golden Ratio (Φ, approximately 1.618). Perez posits that these numbers dictate various aspects of genomic organization, from nucleotide proportions to codon distributions and global densities, creating a system characterized by high accuracy, noise resilience, and mathematical equilibrium.
The Foundation: Chargaff's Rules and the Constant 1
The first of Perez's "3 Genomic Numbers," the constant 1, is deeply rooted in Erwin Chargaff's groundbreaking discoveries concerning DNA composition. Chargaff's first parity rule, a cornerstone of molecular biology, states that in double-stranded DNA, the amount of adenine (A) is approximately equal to thymine (T), and the amount of guanine (G) is approximately equal to cytosine (C). This fundamental observation, often expressed as %A = %T and %G = %C, is crucial for maintaining the structural integrity and complementary nature of the DNA double helix.
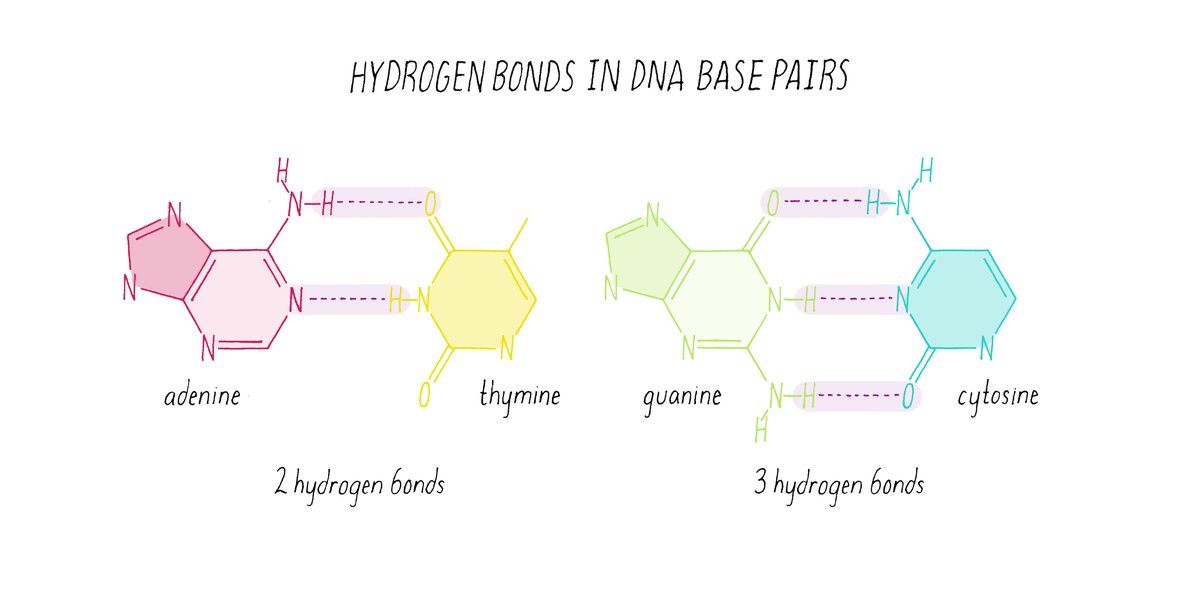
Perez extends this concept beyond simple base pairing, suggesting that the constant 1 represents a global "checksum" mechanism within the genome. This mechanism ensures an optimal balance of atomic mass between DNA strands. The nitrogenous bases have varying atomic masses (A/T are around 135 Da, while G/C are around 151 Da). The parity rule, therefore, contributes to minimizing structural instability by balancing these masses across the strands. Mainstream genomics confirms Chargaff's rules as foundational, noting that they hold asymptotically across genomes due to various structural mechanisms like inversions and transpositions. This aligns with Perez's perspective of the constant 1 acting as a core numerical constraint that guarantees the physical and informational balance of the DNA molecule.
This video provides a clear explanation of Chargaff's rules, which are foundational to understanding the constant 1 in Perez's framework. It elaborates on the base pairing principles (A with T, G with C) and their implications for DNA structure.
The Constant 2: Balancing Codon Frequencies
The second of Perez's proposed genomic numbers, the constant 2, is attributed to the meticulous balancing of codon frequencies within the Universal Genetic Code (UGC). The UGC is the set of rules by which information encoded in genetic material (DNA or RNA sequences) is translated into proteins by living cells. It comprises 64 possible triplet combinations (codons) that encode 20 amino acids and stop signals, exhibiting a characteristic known as degeneracy—meaning multiple codons can specify the same amino acid.
Perez suggests a "2-fold" harmony within this code, where codons are often grouped into pairs that differ by a single nucleotide. This pairing, he argues, plays a crucial role in optimizing the frequency distributions of codons. In the human genome, these distributions average around 2% per triplet, creating a highly balanced system. This balance contributes to the genome's "fractal-like language," which is remarkably resilient to mutations. By distributing codon usage evenly, the impact of a single nucleotide change is minimized, reducing the potential for error propagation and maintaining genetic stability.

Matrix models of the genetic code, such as those employing Kronecker powers or Hadamard matrices, have been explored by various researchers and are seen to align with Perez's concept. These models reveal algebraic symmetries that inherently encode such balances, indicating that triplet populations are predetermined with a remarkable accuracy of approximately 99.9% in large genomes. This perspective implies a highly organized and mathematically precise mechanism governing the translation of genetic information into proteins.
The Golden Ratio (Φ): Fine-Tuning Global Densities
The third and arguably most intriguing of Perez's "3 Genomic Numbers" is the Golden Ratio, Φ (approximately 1.618). This irrational number, found ubiquitously in nature and art, is posited by Perez to fine-tune global densities across various scales within the human genome. This fine-tuning manifests in codon usage biases, GC-content gradients (the proportion of guanine and cytosine bases), and the fractal repetitions observed in DNA sequences.

Perez suggests that Φ structures the genome's "meta-architecture," where triplet counts and base densities follow Fibonacci-like progressions. This ensures a delicate equilibrium between informational entropy (the measure of randomness or disorder in a system) and physical stability. For instance, Φ ratios can be observed in the distribution of purines (A+G) versus pyrimidines (C+T), which might optimize energy efficiency and enhance noise resistance. This perspective aligns with evolutionary models where the UGC functions as a checksum matrix—a hyper-precise numerical grid that verifies sequence integrity, similar to error-detecting codes in computer science.
The result, according to Perez, is a genome operating as a deterministic program where "Form" (semantic coding) and "Substance" (molecular mass and structure) harmonize mathematically. This mathematical precision leads to genomes complying with these rules with over 99% accuracy. While mainstream biology often attributes such patterns to natural selection pressures, Perez's model introduces the concept of a pre-programmed design, highlighting deterministic elegance. Empirical validation for some aspects comes from analyzing human chromosome data, which has shown Φ-modulated waves in base compositions. Such insights could have significant implications for fields like synthetic biology and error-correcting genomics.
The Genomic Meta-Architecture: A Universal Checksum
Perez's framework describes the Universal Genetic Code not just as a set of rules for protein synthesis, but as a sophisticated "checksum matrix." In computer science, a checksum is a small-sized datum computed from a block of digital data for the purpose of detecting errors that may have been introduced during its transmission or storage. Applied to the genome, this concept implies that the genetic code possesses inherent error-checking capabilities, leveraging the "3 Genomic Numbers" to maintain its integrity.
This meta-architecture is said to predetermine triplet populations with an astonishing 99.9% accuracy. This high degree of predictability suggests a deeply embedded numerical order rather than purely stochastic processes. The constraints imposed by these three numbers optimize the balance of atomic mass between DNA strands, contributing to the genome's resilience against noise and mutations. This resilience is a critical feature for any informational system that must faithfully transmit vast amounts of data across generations.
This mindmap illustrates the core components and interconnections of Jean-Claude Perez's "3 Genomic Numbers" framework, demonstrating how each constant contributes to a proposed meta-architecture for the human genome.
The genome, in this view, operates as a deterministic program where informational "Form" and structural "Substance" achieve a state of perfect mathematical equilibrium. "Form" refers to the semantic coding and genetic information, while "Substance" pertains to the physical properties of DNA, such as atomic mass and molecular structure. This harmonious integration suggests a highly optimized system designed for both robustness and efficient information storage and retrieval.
Perspectives and Consensus
While Perez's "3 Genomic Numbers" framework presents a compelling and mathematically elegant model of genomic organization, it is important to note its position within the broader scientific community. Chargaff's first parity rule (A=T, G=C) is a universally accepted and foundational principle in genomics. Chargaff's second parity rule (PR-2), which suggests an intra-strand symmetry (A≈T and G≈C within a single strand), also has some support in the literature, emerging asymptotically in many genomes, though it remains a subject of ongoing debate.
However, Perez's extensions, particularly the specific roles assigned to the constant 2 and the Golden Ratio (Φ) in predetermining codon frequencies and fine-tuning global densities with 99.9% accuracy, are not widely accepted in mainstream genomics. His claims are primarily published in niche outlets, research blogs, and certain journals that may not be considered central to the field. This means that while the ideas are intriguing, they have not undergone broad independent replication and validation, which are critical steps for any scientific theory to gain widespread acceptance.
This radar chart compares the perceived strengths of Perez's "3 Genomic Numbers" framework with the widely accepted Chargaff's First Parity Rule across several analytical dimensions. It highlights areas where Perez's theory offers strong conceptual integration and elegance, contrasting with the higher mainstream acceptance and empirical evidence for Chargaff's established rule.
Despite the lack of mainstream consensus, the intellectual endeavor behind Perez's work underscores a persistent quest to uncover deeper mathematical principles governing biological systems. Understanding these concepts provides valuable context for how scientific theories evolve and how novel ideas, even if initially outside the mainstream, can stimulate further research and discussion.
The Interplay of Form and Substance
The concept of "Form" and "Substance" achieving perfect mathematical equilibrium is central to Perez's hypothesis. "Form" can be understood as the informational content and coding patterns of the DNA—the sequence of nucleotides that dictates the production of proteins and the regulation of cellular processes. "Substance," on the other hand, refers to the physical and chemical properties of the DNA molecule itself, such as the atomic masses of its constituent bases, its three-dimensional structure, and its energetic stability.
Perez argues that the "3 Genomic Numbers" ensure that these two aspects are not merely coincidentally linked but are intrinsically and mathematically intertwined. The balance of atomic masses (driven by the constant 1), the optimized codon frequencies (influenced by the constant 2), and the fine-tuned global densities (governed by Φ) collectively create a system where the information encoded within the DNA is seamlessly supported by its physical structure. This intrinsic harmony contributes to the genome's remarkable efficiency, stability, and capacity for precise self-replication.
The Universal Genetic Code as a Deterministic System
The idea of the genome as a "deterministic program" challenges the view that genomic evolution is primarily a result of random mutations filtered by natural selection. Instead, it suggests an inherent, pre-programmed design where outcomes are highly predictable based on underlying mathematical rules. While this perspective is not widely adopted in evolutionary biology, it offers an alternative lens through which to view the complexity and apparent optimization of biological systems.
This deterministic view implies that the very structure of the genetic code, mediated by these numerical constants, guides and constrains evolutionary pathways. It suggests that certain genomic configurations are inherently more stable, efficient, and robust due to their adherence to these mathematical principles. Such a framework could potentially explain the high conservation of certain genomic elements and the recurring patterns observed across diverse species.
This bar chart compares the emphasis on deterministic elements in Perez's framework versus the role of stochasticity in mainstream evolutionary theory across several key aspects of genomic function and evolution. It highlights how Perez's model attributes higher predictability and inherent optimization to the genome's design.
Implications for Future Research
If Perez's "3 Genomic Numbers" framework were to gain broader acceptance, it could open new avenues for research in several areas:
- Synthetic Biology: A deeper mathematical understanding of genomic design principles could inform the construction of novel biological systems with enhanced stability, efficiency, and predictability.
- Error-Correcting Genomics: The concept of the genetic code as a checksum matrix could lead to new methods for detecting and correcting genomic errors, with applications in disease diagnosis and gene therapy.
- Evolutionary Biology: While challenging current paradigms, the framework could stimulate new investigations into the mathematical underpinnings of evolutionary processes and the potential for non-random constraints on genomic change.
- Bioinformatics and Data Analysis: New algorithms and analytical tools could be developed to search for and validate these numerical patterns in vast genomic datasets, potentially uncovering previously unnoticed correlations.
Ultimately, whether Perez's comprehensive framework becomes a widely accepted model or remains a provocative alternative, it contributes to the ongoing scientific discourse about the fundamental nature of life and the intricate beauty of its genetic code.
Frequently Asked Questions
Conclusion
Jean-Claude Perez's "3 Genomic Numbers" theory offers a unique and mathematically driven perspective on the organization of the human genome. By positing that the constants 1, 2, and the Golden Ratio (Φ) govern fundamental aspects of DNA, from base parity and codon frequencies to global densities, the framework suggests a self-designed, deterministic system operating with remarkable precision. While building upon the established foundation of Chargaff's rules, Perez's extensions propose a meta-architecture where the Universal Genetic Code functions as a checksum matrix, ensuring high accuracy and noise resilience. This vision of the genome as a program in perfect mathematical equilibrium between informational "Form" and structural "Substance" is both intellectually stimulating and thought-provoking, even as it navigates the complex terrain between established scientific consensus and emerging alternative theories.
Recommended Further Exploration
- How do Chargaff's rules impact DNA stability?
- What are the mathematical properties of the Universal Genetic Code?
- Can the Golden Ratio be found in other biological systems?
- What are the implications of a deterministic genome for evolutionary theory?