
Decoding the Symphony of Gene Expression: Nascent RNA and Transcriptional Pausing
Unraveling the intricate dance between newly synthesized RNA and the temporary halts of RNA polymerase, crucial for precise gene regulation.
Key Insights into the Interplay
- Dynamic Regulation: Transcriptional pausing is an evolutionarily conserved mechanism that temporarily halts RNA polymerase (RNAP) during RNA synthesis, acting as a crucial checkpoint for gene expression.
- Real-time Monitoring: Nascent RNA represents the newly synthesized RNA molecules still associated with the transcription machinery, offering a real-time snapshot of active gene expression and immediate regulatory changes.
- Coordinated Events: The interplay between nascent RNA and transcriptional pausing allows for the precise coordination of co-transcriptional processes like RNA folding, splicing, and the recruitment of regulatory proteins, ensuring proper gene biogenesis and cellular response.
In the complex world of molecular biology, the synthesis of RNA from a DNA template—a process known as transcription—is meticulously controlled. Two fundamental concepts, nascent RNA and transcriptional pausing, are intimately connected and play pivotal roles in orchestrating gene expression, RNA biogenesis, and a cell's response to various cues. This exploration delves into these concepts, highlighting their mechanisms, significance, and intricate relationship.
Understanding Nascent RNA: The Real-Time Transcriptional Snapshot
Capturing the immediate output of genetic information.
Nascent RNA refers to the RNA molecules that are actively being synthesized by RNA polymerase or have just been synthesized and remain associated with the transcription complex or chromatin. Unlike mature RNA, which has undergone extensive processing like splicing, capping, and polyadenylation, nascent RNA provides a direct, dynamic, and real-time readout of transcriptional activity. It represents the raw, unfiltered output of gene expression, making it invaluable for studying immediate cellular responses.
The analysis of nascent RNA offers several distinct advantages over measuring steady-state RNA levels:
- Immediate Regulatory Changes: Nascent RNA analyses can reveal rapid transcriptional responses to perturbations that might be obscured in steady-state measurements due to RNA degradation or processing.
- Detection of Unstable Transcripts: It allows for the capture of short-lived and unstable transcripts, such as enhancer RNAs (eRNAs) and upstream antisense RNAs (uaRNAs), which are often challenging to detect with traditional RNA sequencing methods. eRNAs, in particular, serve as crucial markers of active enhancers.
- Insights into Enhancer Activity and Gene Regulation: Technologies like Global Nuclear Run-On sequencing (GRO-seq) and Precision Run-On sequencing (PRO-seq) enable simultaneous quantification of transcription in enhancers and their target genes. This provides a comprehensive view of enhancer-mediated gene regulation, facilitating the annotation and quantification of active enhancers and the prediction of their target genes.
- Understanding Transcription Dynamics: Nascent RNA analysis provides insights into the dynamics of transcriptional processing and regulation at multiple stages, including RNA polymerase recruitment, transcription initiation, elongation, and termination. Single-cell nascent RNA sequencing (e.g., scGRO-seq) further allows for deciphering transcription dynamics and enhancer-gene relationships at the individual cell level.
Various complementary methods, including live-cell imaging, metabolic labeling of RNA, and RNA-seq of purified nascent RNA, have been developed to quantitatively track nascent transcription genome-wide at nucleotide resolution. These techniques continue to provide novel insights into the mechanisms of gene regulation and transcription-coupled RNA processing.
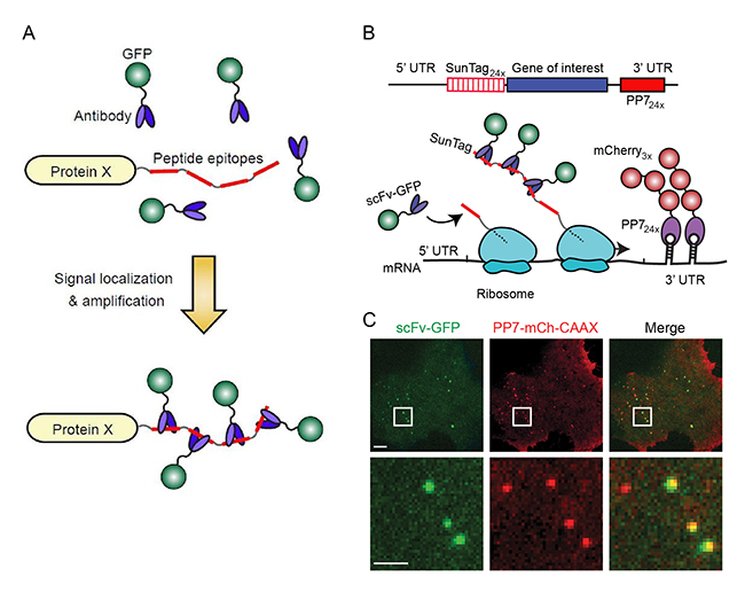
Visual representation of nascent RNA synthesis and its association with cellular structures, providing a snapshot of active gene expression.
The Mechanism of Transcriptional Pausing: A Strategic Halt in Synthesis
Orchestrating precision through temporary arrests.
Transcriptional pausing refers to a transient halt of RNA polymerase during the elongation phase of transcription, where the polymerase temporarily stops RNA synthesis after transcribing a short region of DNA, typically 20-60 nucleotides downstream of the transcription start site. This evolutionarily conserved mechanism plays a crucial role in regulating gene expression across all domains of life, from bacteria to mammals. These pauses are not random; they are induced by specific signals encoded in the nucleic acid sequence (DNA and RNA sequences and structures) and allow time for various co-transcriptional events to occur.
Classes of Transcriptional Pause Signals:
There are four recognized classes of transcriptional pause signals:
- Elemental Pauses: These represent an elemental paused state from which longer-lived pauses can arise. The RNAP rearranges into a catalytically inactive conformation, unable to load the next nucleotide triphosphate (NTP) substrate.
- Backtrack Pauses: These pauses are often stabilized when the RNA-DNA hybrid is destabilized, such as by a UA-rich RNA-DNA hybrid or nucleotide misincorporation. Backtracking involves the RNAP moving backward along the DNA template, disengaging the nascent RNA from the active site. This is often linked to error correction or regulatory delays.
- Hairpin-Stabilized Pauses: These pauses are often associated with the formation of secondary structures (hairpins) in the nascent RNA, which can cause an allosteric conformational change in the RNAP, delaying translocation and misalignment.
- Regulator-Stabilized Pauses: These pauses are influenced by the binding of transcription regulatory proteins or complexes that interact with the elongation complex to stabilize pausing. For example, in metazoans, negative elongation factor (NELF) and DRB sensitivity inducing factor (DSIF) mediate promoter-proximal pausing of RNA Polymerase II (Pol II).
Functional Significance of Pausing:
Transcriptional pausing serves several critical functions:
- Coordination of Co-transcriptional Events: Pauses create windows of time and space for other regulatory mechanisms to take place, such as RNA folding, splicing, and interactions with cellular molecules and complexes like ribosomes. In many organisms, including yeast, splicing often occurs cotranscriptionally, meaning it happens while the RNA is still being transcribed. Pausing near the end of genes provides sufficient time for splicing to occur before transcription termination.
- Regulation of Productive Elongation: Pol II pausing regulates the productive elongation step of transcription at key genes, responding to various signaling pathways.
- Control of Gene Expression Dynamics: The rate and duration of pausing are sequence-dependent and can affect a gene's transcriptional dynamics, contributing to noise in gene expression and phenotypic diversity.
- Signal Integration: Pausing enables the integration of various signals and fine-tuning of transcriptional responses. Mutations in regulators of pause-release have been linked to neurodevelopmental disorders, highlighting their importance.
- Setting Up Poised Transcriptional States: Pausing can prepare genes for rapid activation, allowing promoters to be "poised" for immediate activation in response to signals.
- Ensuring Transcription Fidelity: Pauses allow for the correction of misincorporated nucleotides or facilitate backtracking for proofreading.
This radar chart illustrates the perceived importance and analytical power of transcriptional pausing and nascent RNA across various aspects of gene regulation. It highlights how these two concepts contribute uniquely and synergistically to cellular processes.
The Interconnection: How Nascent RNA Illuminates Pausing
A real-time window into the kinetic events of transcription.
The study of nascent RNA is crucial for understanding transcriptional pausing because nascent RNA sequencing directly measures the presence and density of actively engaged RNA polymerases along the genome. This allows researchers to pinpoint where and for how long RNAP pauses. Transcriptional pausing directly influences the production and characteristics of nascent RNA, creating a regulatory feedback loop in gene expression.
When RNAP pauses, it limits the frequency and speed of transcription initiation, which in turn affects how nascent RNA is synthesized and processed. Pauses can extend the duration of nascent RNA association with the transcription complex, allowing time for regulatory events like RNA folding or protein binding. By analyzing nascent RNA, scientists can:
- Identify promoter-proximal pausing events.
- Quantify the duration of pauses.
- Investigate how pausing restricts transcriptional output at different gene classes.
- Determine the impact of regulators on pause entry and exit.
Nascent RNA sequencing studies reveal that pauses occur at key genomic regions, such as enhancers and promoters, coordinating the burst-like production of nascent transcripts. This deep insight into the dynamics of transcriptional processing and regulation at multiple stages—including RNA polymerase recruitment, transcription initiation, elongation, and termination—is fundamental to understanding gene regulation.
This video, "Mike Levine (UC Berkeley) Part 3: Transcriptional Precision: Paused Polymerase II...", provides a detailed academic perspective on the role of paused polymerase II in ensuring transcriptional precision. It highlights how these temporary halts are critical for fine-tuning gene expression, offering an in-depth look at a key aspect of transcriptional pausing.
Key Technologies and Their Contributions
Advancements in measuring the real-time dynamics of transcription.
Modern biological research has been revolutionized by technologies capable of probing nascent RNA and transcriptional pausing with unprecedented resolution. These techniques provide crucial insights into how cells regulate gene activity at the molecular level.
| Technique | Description | Key Insights Provided |
|---|---|---|
| Global Nuclear Run-On Sequencing (GRO-seq) | Measures actively transcribing RNA polymerase across the genome by allowing elongation in vitro with labeled nucleotides. | Identifies active enhancers and promoters, quantifies transcription rates, and maps RNA polymerase density genome-wide. |
| Precision Run-On Sequencing (PRO-seq) | A higher resolution variant of GRO-seq that maps the exact position of RNA polymerase and nascent RNA 3' ends. | Pinpoints precise transcriptional start sites (TSSs), identifies pause sites with nucleotide resolution, and reveals the directionality of transcription. |
| Single-cell Nascent RNA Sequencing (scGRO-seq) | Adapts nascent RNA profiling to single-cell resolution. | Unveils cell-to-cell variability in gene expression dynamics, identifies enhancer-gene relationships in heterogeneous cell populations, and deciphers transcription dynamics at the individual cell level. |
| Metabolic Labeling of RNA | Incorporates modified nucleotides (e.g., 4-thiouridine) into newly synthesized RNA, which can then be isolated. | Measures RNA synthesis rates and turnover, identifies unstable transcripts, and provides temporal resolution of gene expression changes. |
| Live-cell Imaging | Uses fluorescent reporters to visualize transcription events in real-time within living cells. | Observes transcription bursts, tracks RNA polymerase movement, and visualizes the dynamics of gene activation and repression. |
This table summarizes prominent techniques used to study nascent RNA and transcriptional pausing, highlighting their unique capabilities in revealing the intricate dynamics of gene expression.
The Broader Biological Significance
Impacts on development, disease, and cellular adaptation.
The precise control exerted by nascent RNA dynamics and transcriptional pausing is fundamental to life. This regulatory layer ensures that genes are expressed at the right time, in the right amount, and in the right cellular context.
- Development and Differentiation: Pausing is critical during development and differentiation to ensure precise temporal gene expression, guiding cell fate decisions and tissue formation. For instance, in sprouting angiogenesis, pausing modulates Pol II activity to fine-tune nascent RNA output, essential for proper vascular development.
- Disease Implications: Alterations or dysregulation in the pausing machinery have been implicated in various diseases, including neurodevelopmental disorders and cancer. Understanding these mechanisms could lead to new therapeutic strategies.
- Cellular Adaptation: Pausing mechanisms allow cells to integrate environmental cues, stress signals, or developmental instructions by dynamically modulating transcriptional output. This adaptability is crucial for survival and response to external challenges.
- Evolutionary Conservation: The presence of transcriptional pausing in diverse organisms, from bacteria to plants and humans, underscores its fundamental importance as a conserved mechanism of genetic regulation. While the core principles are shared, specific features of nascent RNA and pausing may differ, reflecting diverse regulatory strategies adapted to different biological contexts.
Immediate transcriptional output"] id4["Analytical Significance: Real-time gene expression
Detects unstable transcripts (eRNAs)"] id5["Techniques: GRO-seq
PRO-seq
scGRO-seq
Live-cell Imaging"] id6["Transcriptional Pausing"] id7["Definition: Temporary halt of RNAP"] id8["Mechanisms: Elemental pauses
Backtrack pauses
Hairpin-stabilized pauses
Regulator-stabilized pauses"] id9["Regulatory Roles: Coordinates co-transcriptional events (splicing, folding)
Integrates signals
Regulates elongation
Ensures fidelity"] id10["Factors: NELF
DSIF"] id11["Interconnection"] id12["Nascent RNA measures RNAP density & pause duration"] id13["Pausing limits transcription initiation frequency"] id14["Pauses create 'windows of time and space' for processing"] id15["Affects nascent RNA production patterns"] id16["Biological Significance"] id17["Crucial for Gene Regulation"] id18["Impacts Development & Differentiation"] id19["Linked to Neurodevelopmental Disorders"] id20["Facilitates Cellular Adaptation to Stress"]
This mindmap visually outlines the intricate relationship between nascent RNA and transcriptional pausing, illustrating their definitions, mechanisms, interconnections, and broader biological significance. It serves as a comprehensive overview of the key concepts discussed.
Frequently Asked Questions (FAQ)
Conclusion
Nascent RNA and transcriptional pausing are not isolated phenomena but rather deeply intertwined processes that form a critical regulatory layer in gene expression. Transcriptional pausing acts as a sophisticated molecular brake, ensuring that RNA synthesis is not a mere linear progression but a carefully choreographed sequence of events. This pausing allows for the real-time integration of cellular signals, the coordination of complex co-transcriptional processes like splicing, and the overall fine-tuning of gene output. Concurrently, nascent RNA, as the immediate product of this dynamic transcription, provides the empirical evidence for these kinetic events, offering an unparalleled window into the immediate regulatory responses of a cell. The synergistic study of nascent RNA and transcriptional pausing, empowered by advanced sequencing and imaging technologies, continues to unravel profound insights into fundamental biological principles, with far-reaching implications for understanding development, disease, and cellular adaptation.
Recommended Further Exploration
- Explore the molecular mechanisms of RNA polymerase backtracking and its role in transcriptional fidelity.
- Delve deeper into the fascinating role of enhancer RNAs (eRNAs) in enhancer-mediated gene regulation.
- Investigate the cutting-edge applications of single-cell nascent RNA sequencing in dissecting cellular heterogeneity.
- Understand the clinical relevance of transcriptional pausing in the context of neurodevelopmental disorders.